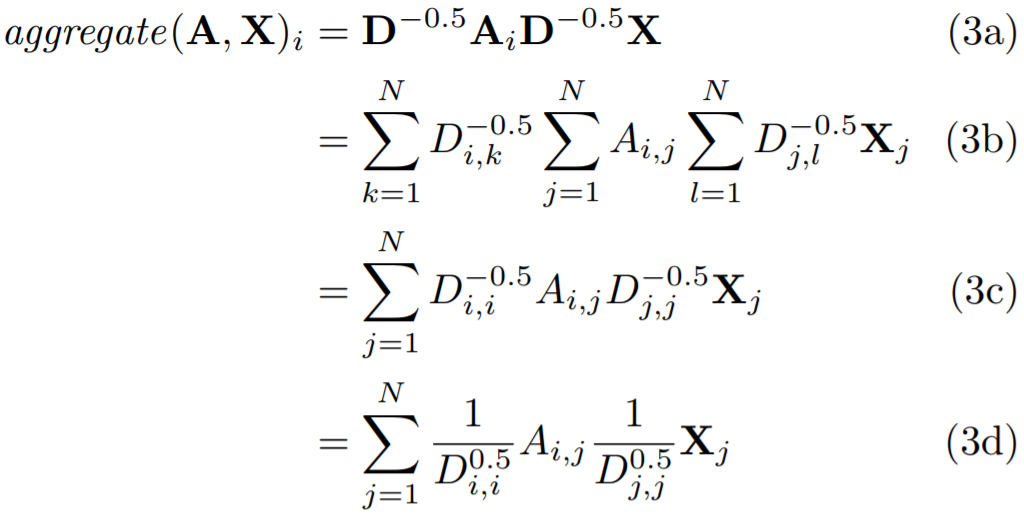#### Loading Required Libraries ####
import torch
import torch.nn as nn
import torch.optim as optim
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
# get_ipython().run_line_magic('matplotlib', 'notebook')
# import imageio
# from celluloid import Camera
# from IPython.display import HTML
# plt.rcParams['animation.ffmpeg_path'] = '/usr/local/bin/ffmpeg'
#### The Convolutional Layer ####
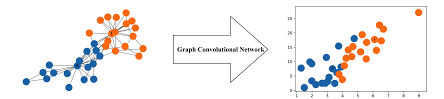
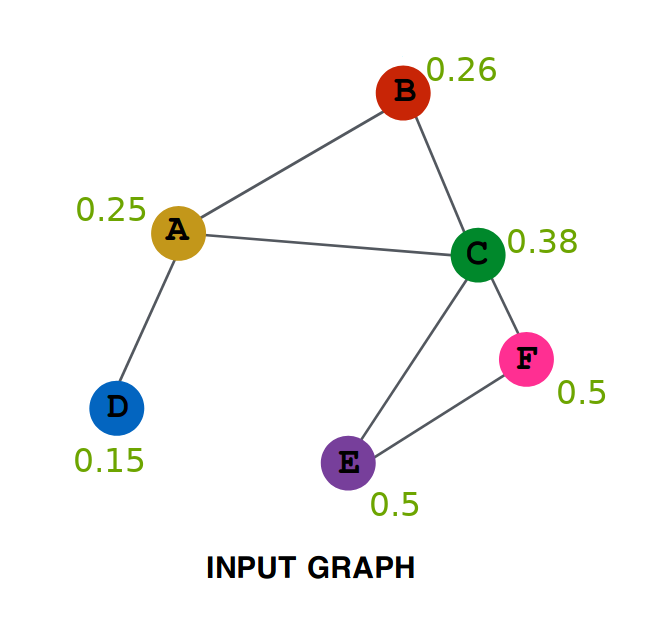
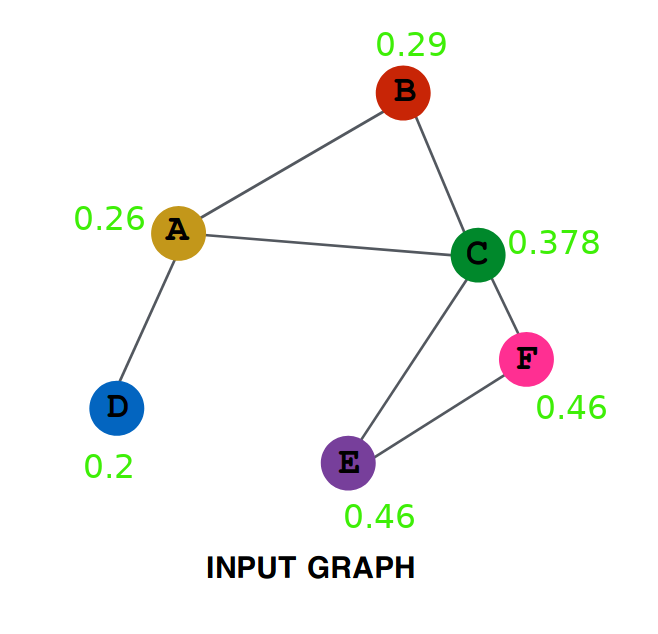
# First we will be creating the GCNConv class, which will serve as the Layer creation class.
# Every instance of this class will be getting Adjacency Matrix as input and will be outputing
# 'RELU(A_hat * X * W)', which the Net class will use.
class GCNConv(nn.Module):
def __init__(self, A, in_channels, out_channels):
super(GCNConv, self).__init__()
self.A_hat = A+torch.eye(A.size(0))
self.D = torch.diag(torch.sum(self.A_hat,1))
self.D = self.D.inverse().sqrt()
self.A_hat = torch.mm(torch.mm(self.D, self.A_hat), self.D)
self.W = nn.Parameter(torch.rand(in_channels,out_channels, requires_grad=True))
def forward(self, X):
out = torch.relu(torch.mm(torch.mm(self.A_hat, X), self.W))
return out
class Net(torch.nn.Module):
def __init__(self,A, nfeat, nhid, nout):
super(Net, self).__init__()
self.conv1 = GCNConv(A,nfeat, nhid)
self.conv2 = GCNConv(A,nhid, nout)
def forward(self,X):
H = self.conv1(X)
H2 = self.conv2(H)
return H2
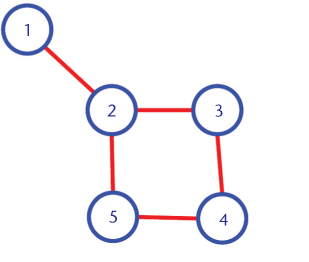
# 'A' is the adjacency matrix, it contains 1 at a position (i,j), if there is a edge between the node i and node j.
A=torch.Tensor([[0,1,1,1,1,1,1,1,1,0,1,1,1,1,0,0,0,1,0,1,0,1,0,0,0,0,0,0,0,0,0,1,0,0],
[1,0,1,1,0,0,0,1,0,0,0,0,0,1,0,0,0,1,0,1,0,1,0,0,0,0,0,0,0,0,1,0,0,0],
[1,1,0,1,0,0,0,1,1,1,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,1,0],
[1,1,1,0,0,0,0,1,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,0,0,0,0,0,1,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,0,0,0,0,0,1,0,0,0,1,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,1,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,1,1],
[0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1],
[1,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,1,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,1,0,1,0,0,1,1],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,1,0,0,0,1,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,1,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,1],
[0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,1],
[0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,1],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,1,0,0,0,0,0,1,1],
[0,1,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,1,0,0,0,1,1],
[0,0,1,0,0,0,0,0,1,0,0,0,0,0,1,1,0,0,1,0,1,0,1,1,0,0,0,0,0,1,1,1,0,1],
[0,0,0,0,0,0,0,0,1,1,0,0,0,1,1,1,0,0,1,1,1,0,1,1,0,0,1,1,1,1,1,1,1,0]
])
# actual label
label = [0, 0, 0, 0 ,0 ,0 ,0, 0, 1, 1, 0 ,0, 0, 0, 1 ,1 ,0 ,0 ,1, 0, 1, 0 ,1 ,1, 1, 1, 1 ,1 ,1, 1, 1, 1, 1, 1 ]
# label for admin(node 1) and instructor(node 34) so only these two contain the known class label (0 and 1)
# all others are set to -1, meaning predicted value of these nodes is ignored in the loss function.
target=torch.tensor([0,-1,-1,-1, -1, -1, -1, -1,-1,-1,-1,-1, -1, -1, -1, -1,-1,-1,-1,-1, -1, -1, -1, -1,-1,-1,-1,-1, -1, -1, -1, -1,-1,1])
# X is the feature matrix.
# Using the one-hot encoding corresponding to the index of the node.
X=torch.eye(A.size(0))
# Network with 10 features in the hidden layer and 2 in output layer.
T=Net(A, X.size(0), 10, 2)
#### Training ####
criterion = torch.nn.CrossEntropyLoss(ignore_index=-1)
optimizer = optim.SGD(T.parameters(), lr=0.01, momentum=0.9)
loss=criterion(T(X),target)
#### Plot animation using celluloid ####
# fig = plt.figure()
# camera = Camera(fig)
# We will train for 200 steps and print the cross entropy loss at every 20 steps. The smaller loss means better performance of the model.
# Note that the model is trained to minimise the cross entropy loss.
for i in range(200):
optimizer.zero_grad()
loss=criterion(T(X), target)
loss.backward()
optimizer.step()
l=(T(X));
# plt.scatter(l.detach().numpy()[:,0],l.detach().numpy()[:,1],c=[0, 0, 0, 0 ,0 ,0 ,0, 0, 1, 1, 0 ,0, 0, 0, 1 ,1 ,0 ,0 ,1, 0, 1, 0 ,1 ,1, 1, 1, 1 ,1 ,1, 1, 1, 1, 1, 1 ])
# for i in range(l.shape[0]):
# text_plot = plt.text(l[i,0], l[i,1], str(i+1))
# camera.snap()
if i%20==0:
print("Cross Entropy Loss: =", loss.item())
# print the predictive accuracy by comparing the predicted label with the actual label for all the nodes.
pred_label = torch.argmax(l, dim=1)
accuracy = accuracy_score(label, pred_label.detach().cpu().numpy())
print("Accuracy: ", accuracy)
# animation = camera.animate(blit=False, interval=150)
# animation.save('./train_karate_animation.mp4', writer='ffmpeg', fps=60)
# HTML(animation.to_html5_video())