Tree-Based Methods
Tree, Forest, and Gradient Boosting
Huanfa Chen - huanfa.chen@ucl.ac.uk
13/12/2025
Last week
- Analysis workflow of supervised learning
- Model evaluation methods: train-test split, cross validation (and extensions)
Objectives of this week
- Understand decision trees and tree-based ensemble methods (random forests, gradient boosting)
- Can apply tree-based methods to real-world problems
Decision Trees
Decision Trees (DT)
- Many types of decision trees: classification and regression trees (CART), C4.5, ID3, CHAID, etc.
- Most trees (except CART) are rarely used these days, and CART is still popular.
- Focus on CART: binary trees for classification and regression
- A CART is a regression tree when the target variable Y is continous, and a classification tree when Y is categorical.
- It is not dependent on type of features (X).
Why DT?
- Non-linear, strong models for classification and regression
- Minimal preprocessing: scale and distributions matter little
- Good interpretation: small trees can be explained
Decision Trees for Classification
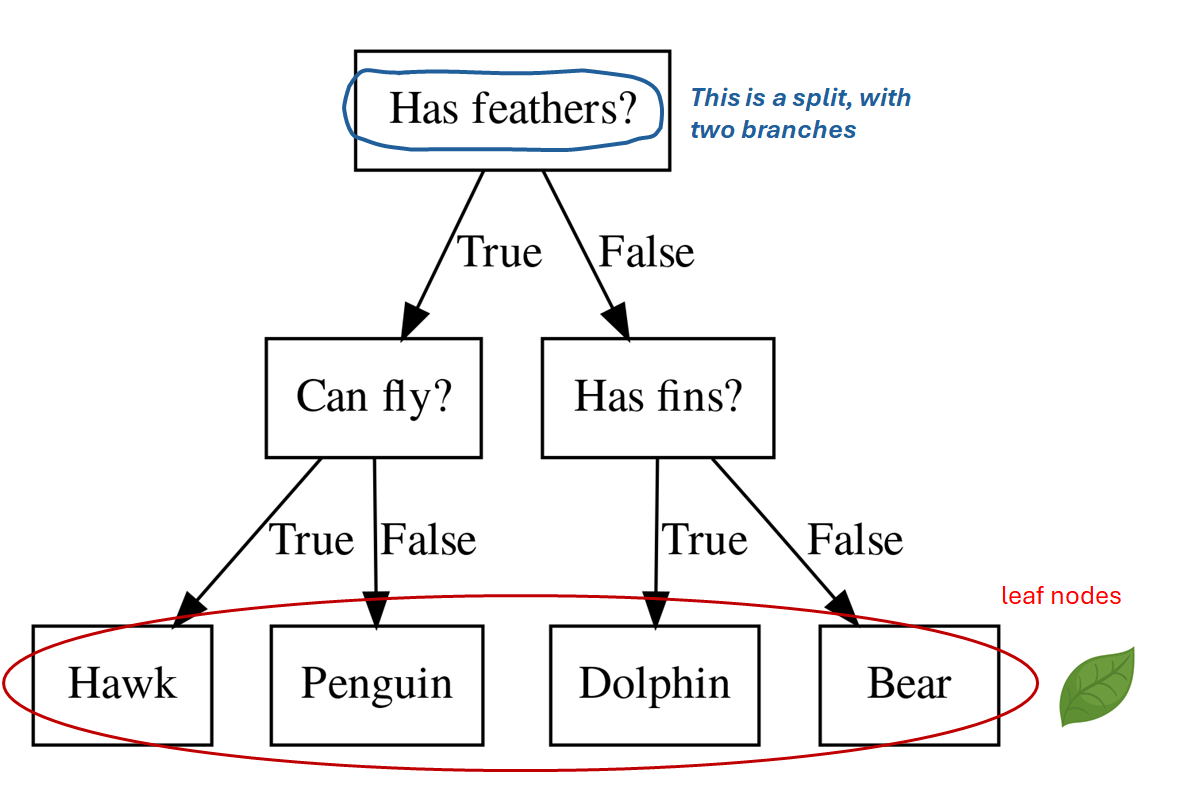
- It starts with a root node
- A split with a threshold on a single X feature divides data into two parts
- Leave Nodes are the endpoints that give predictions.
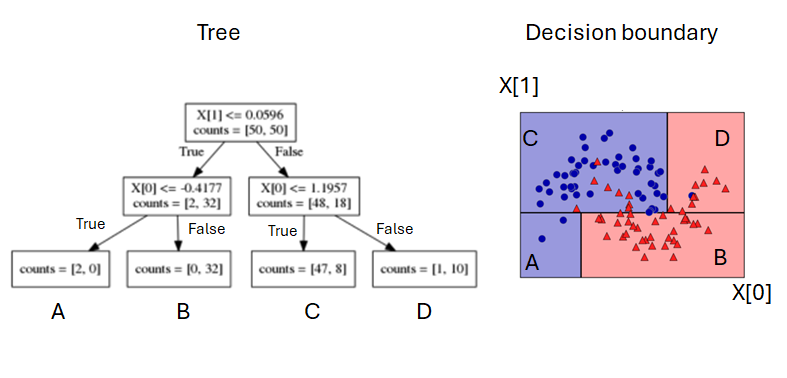
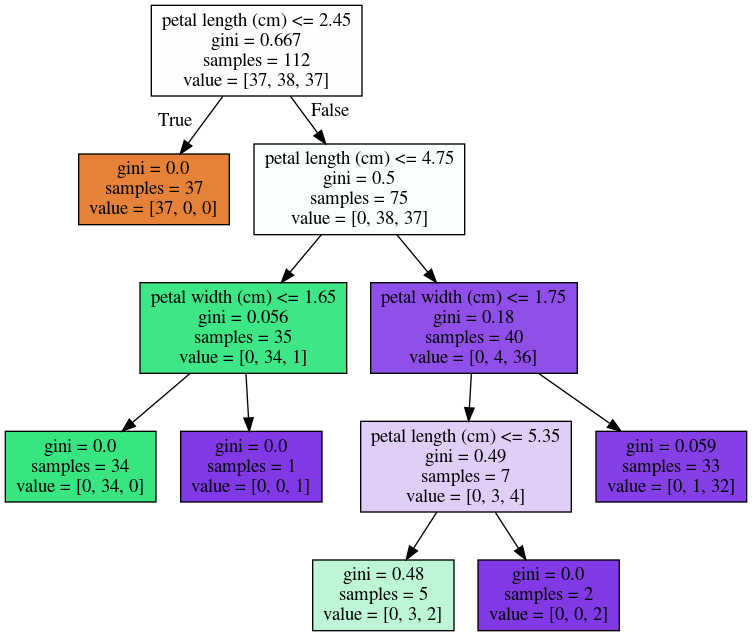
Another (synthetic) classification tree
- A dataset with two numertical features (X[0], [1]), to predict Y with two classes (RED and BLUE)
- Two visuals: tree structure, decision boundary
- The leaf node and region are one-to-one mapped.
- a node with counts=[40, 30] means this node contains 40 RED samples, 30 BLUE samples.

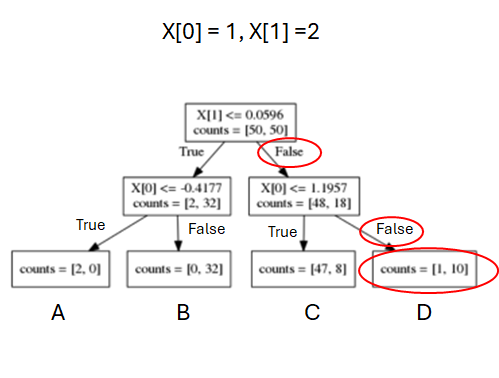
Making predictions with classification tree
- Given a new sample, start from the root node, traverse the tree according to split, until reaching a leaf node.
- Leaf node D has counts=[1, 10], so it predicts BLUE class AND probability distribution of [1/11, 10/11] over two classes.
How to build a classification tree
- A tree is built by a split from the root node, then recursively splitting child nodes.
- A key question: How to choose a split?
- Note:
- A split is a X feature and a threshold.
- Can’t split on Y, as Y is unknown for new datasets.
- Can’t split on multiple X variables.
- A split is chosen to best separate Y classes, by measuring impurity reduction before & after split
To measure impurity of a node
- Gini: \(H_{\text{gini}}(X_m) = \sum_{k \in \mathcal{Y}} p_{mk}(1-p_{mk})\)
- \(p_{mk}\) is class \(k\) proportion in node \(m\)
- The larger Gini, the more impure the node
- When Gini=0, the node is pure (all samples in one class)
- Example: a node with 10 samples in three classes [2,5,3].
- Gini = 0.2 × 0.8 + 0.5 × 0.5 + 0.3 × 0.7 = 0.66
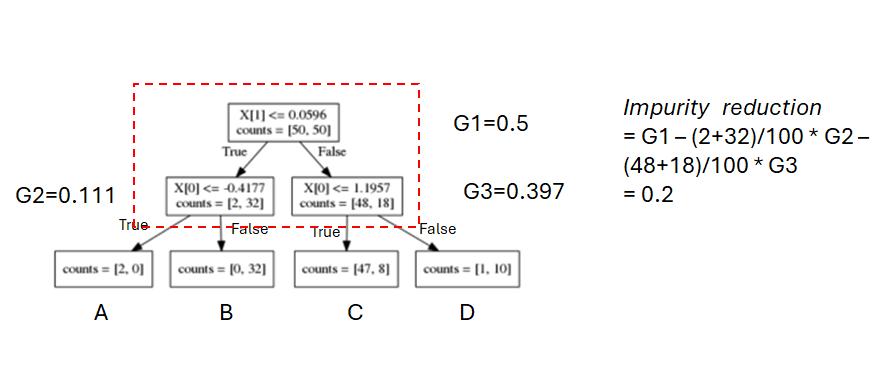
To measure impurity reduction by a split
- A good split will put similar samples together, or generate two child nodes with lower impurity
- Impurity reduction by a split = impurity(parent) - weighted impurity(children)
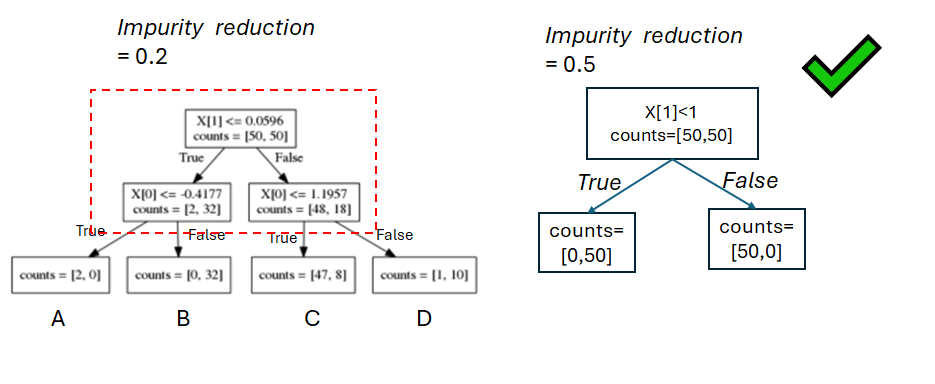
To choose a split
- All splits will be evaluated
- The split with the largest impurity reduction will be chosen
A regression tree
- Similar to classification tree, but a leaf node stores a range of Y values and will predict the mean of Y
- To measure impurity of a node in regresiion tree: Mean Square Error
- \(H(X_m) = \frac{1}{N_m}\sum_{i \in N_m}(y_i-\bar{y}_m)^2\)
- MSE measures the deviation of Y values from its mean, which is similar to the idea of impurity.
- If all Y values are the same, MSE=0, meaning the node is pure.
Summary of impurity measures of decision trees
- For classification:
- Gini (by default)
- Cross-Entropy
- For regression
- Mean Square Error (by default)
- Mean Absolute Error
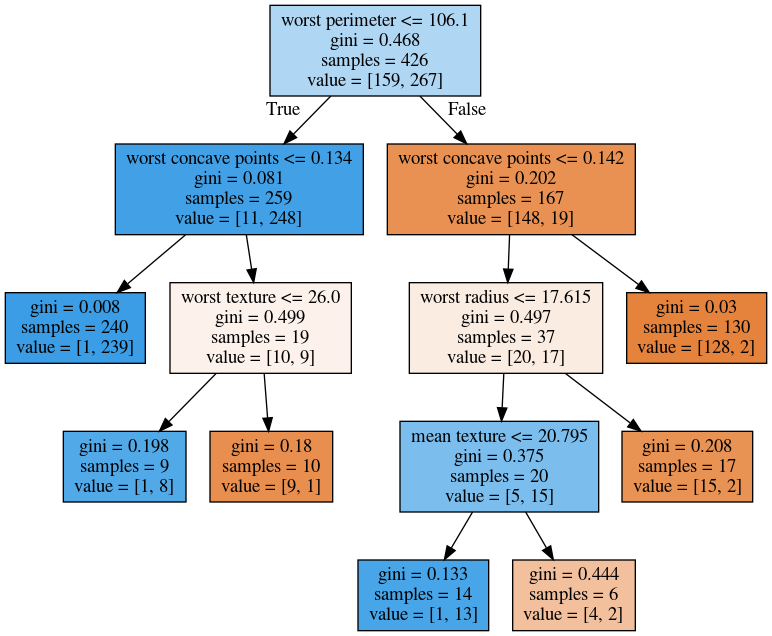
Visualising Trees (sklearn)
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
cancer = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, stratify=cancer.target, random_state=0)
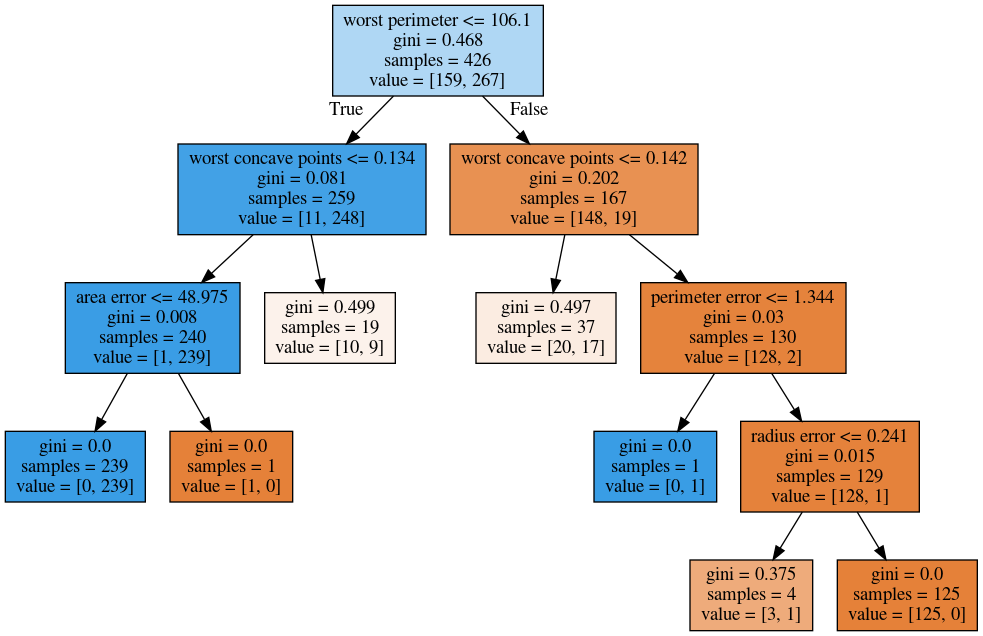
tree = DecisionTreeClassifier(max_depth=2).fit(X_train, y_train)Visualising Trees (plot_tree)
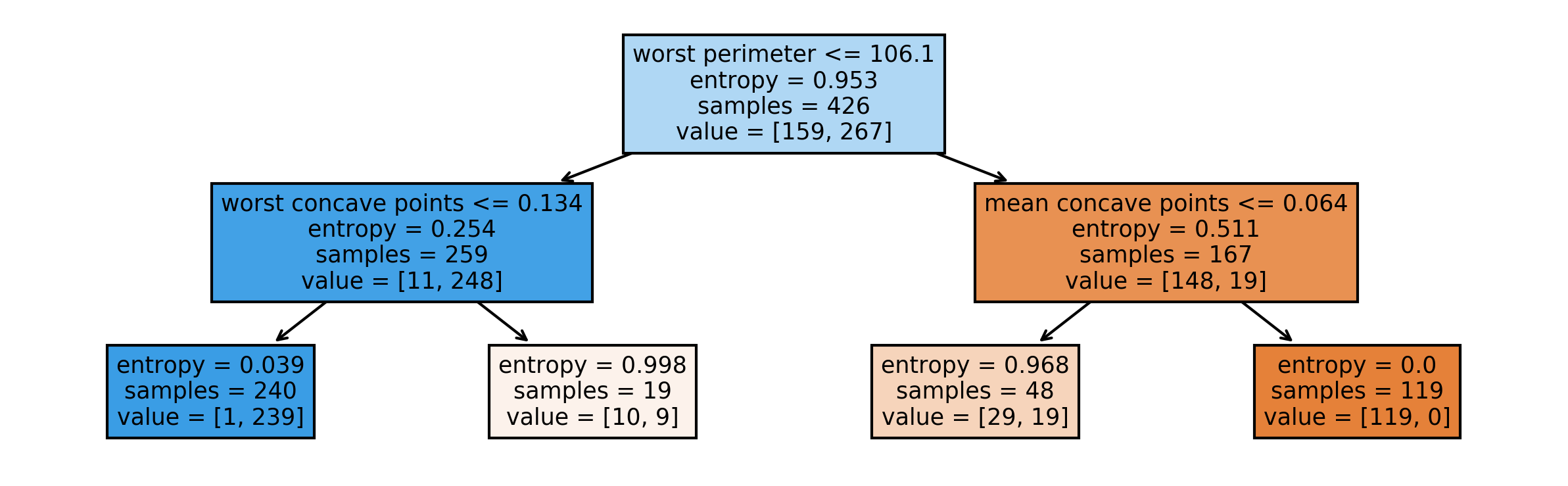
Parameter Tuning
- Pre-pruning (limit growth)
max_depthmax_leaf_nodesmin_samples_splitmin_impurity_decrease
- Post-pruning (cost-complexity) after full growth
No Pruning
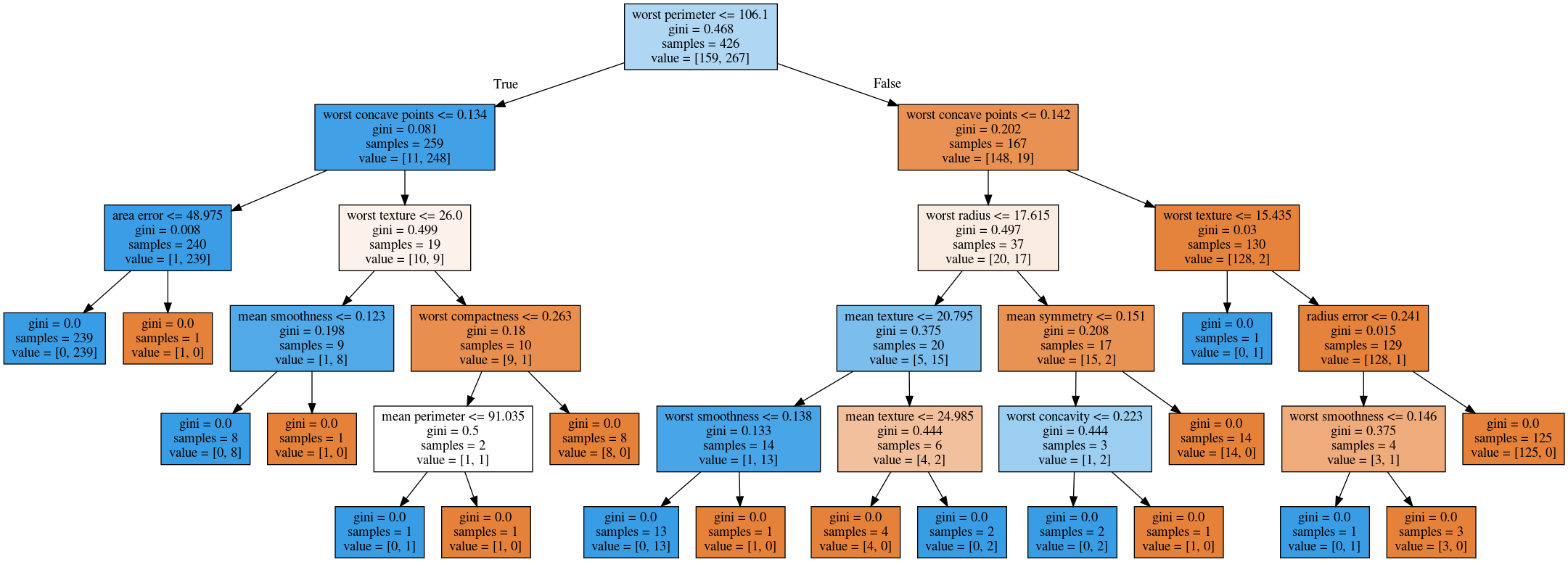
max_depth = 4
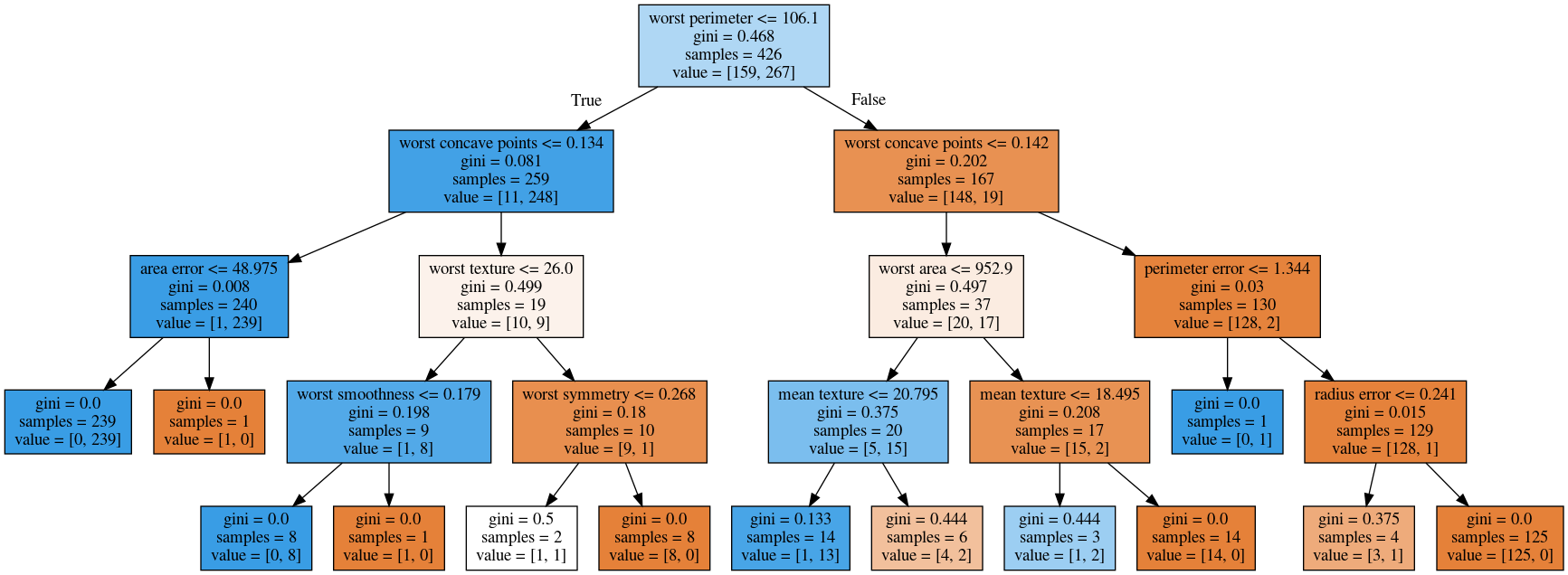
max_leaf_nodes = 8
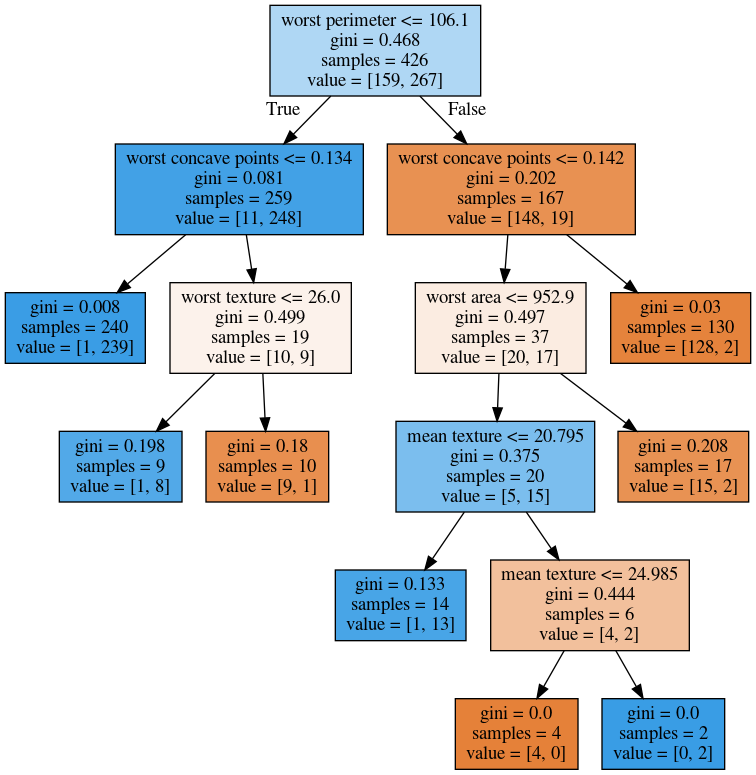
min_samples_split = 50
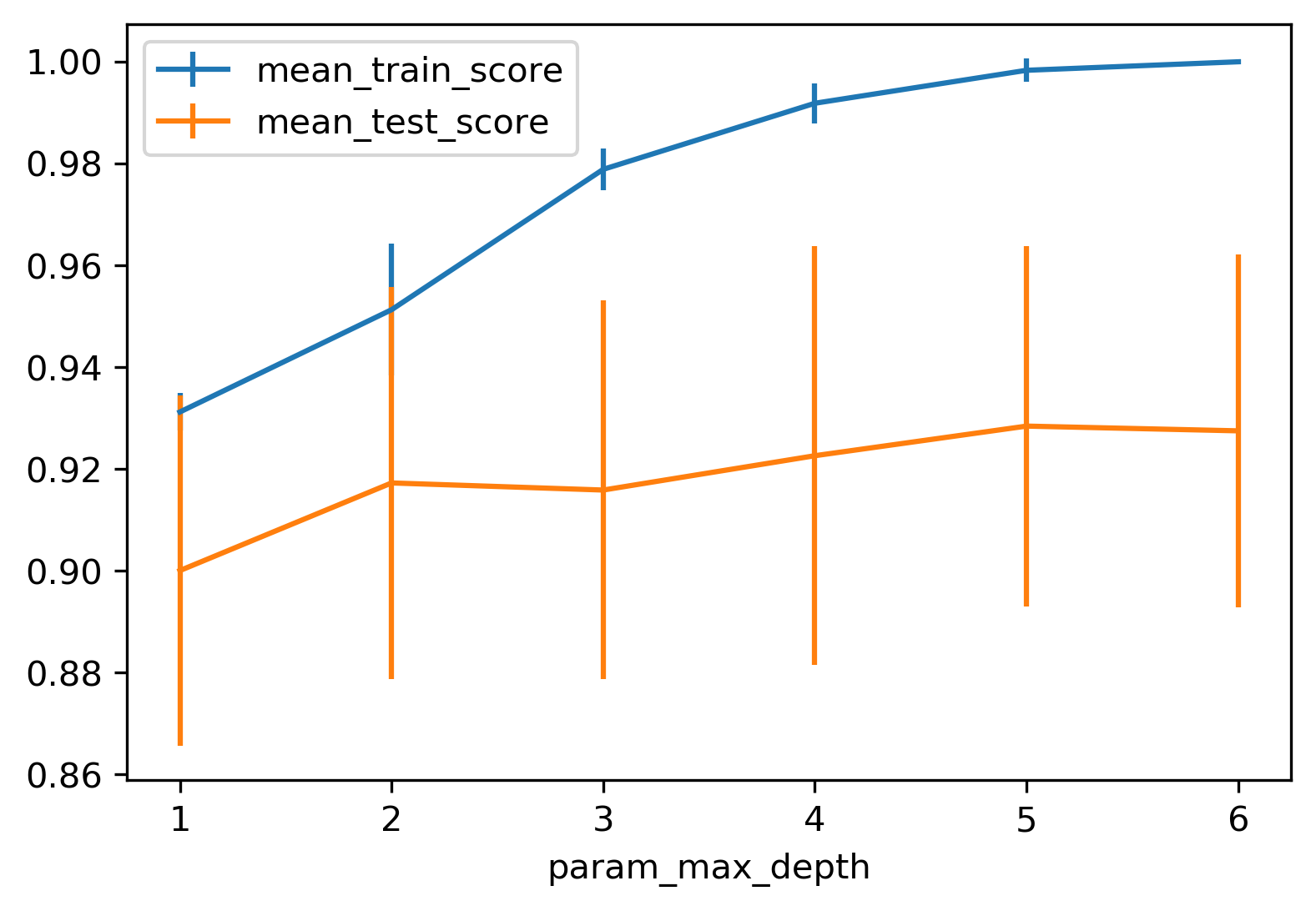
Grid Search: max_depth
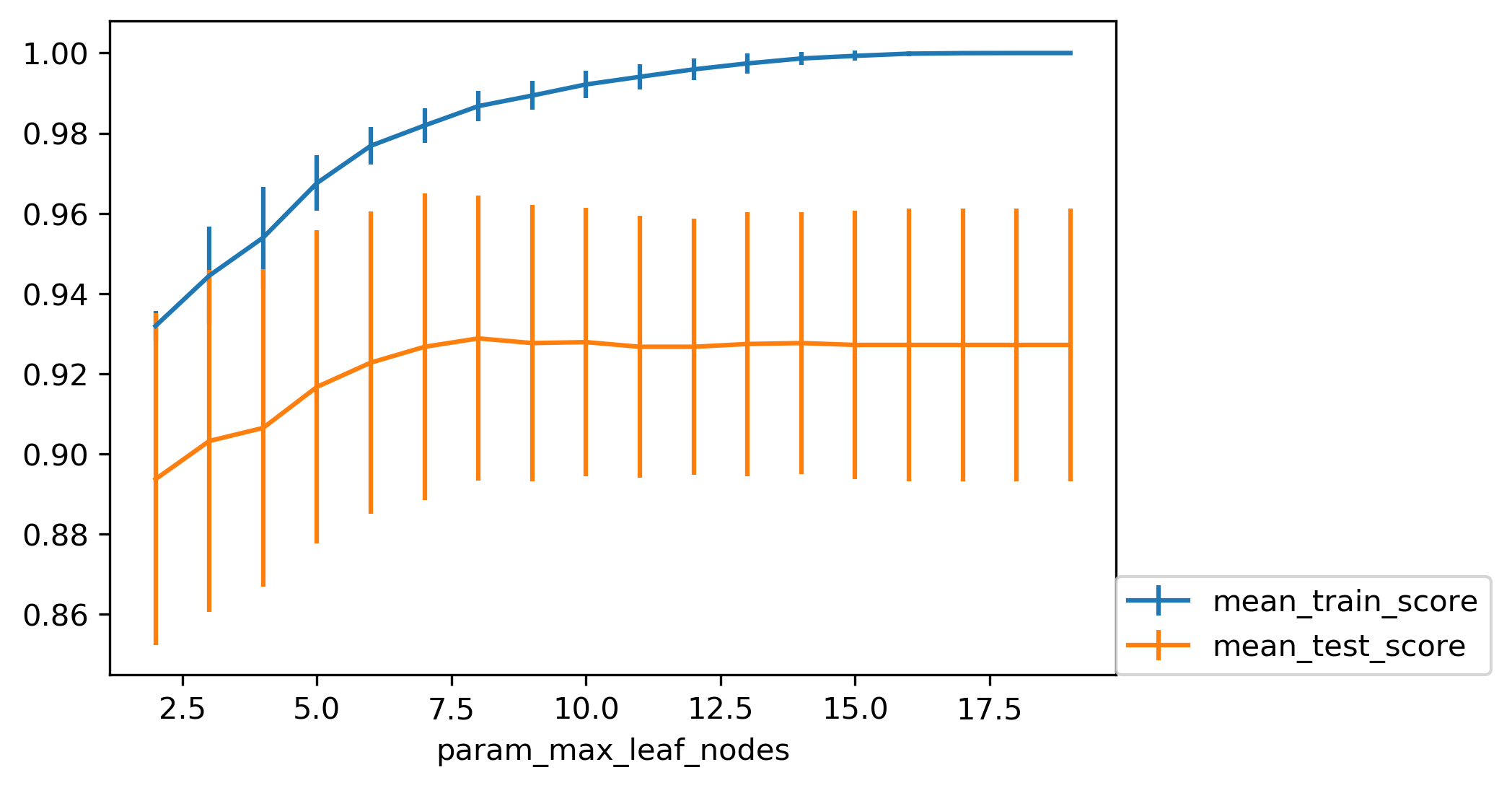
Grid Search: max_leaf_nodes
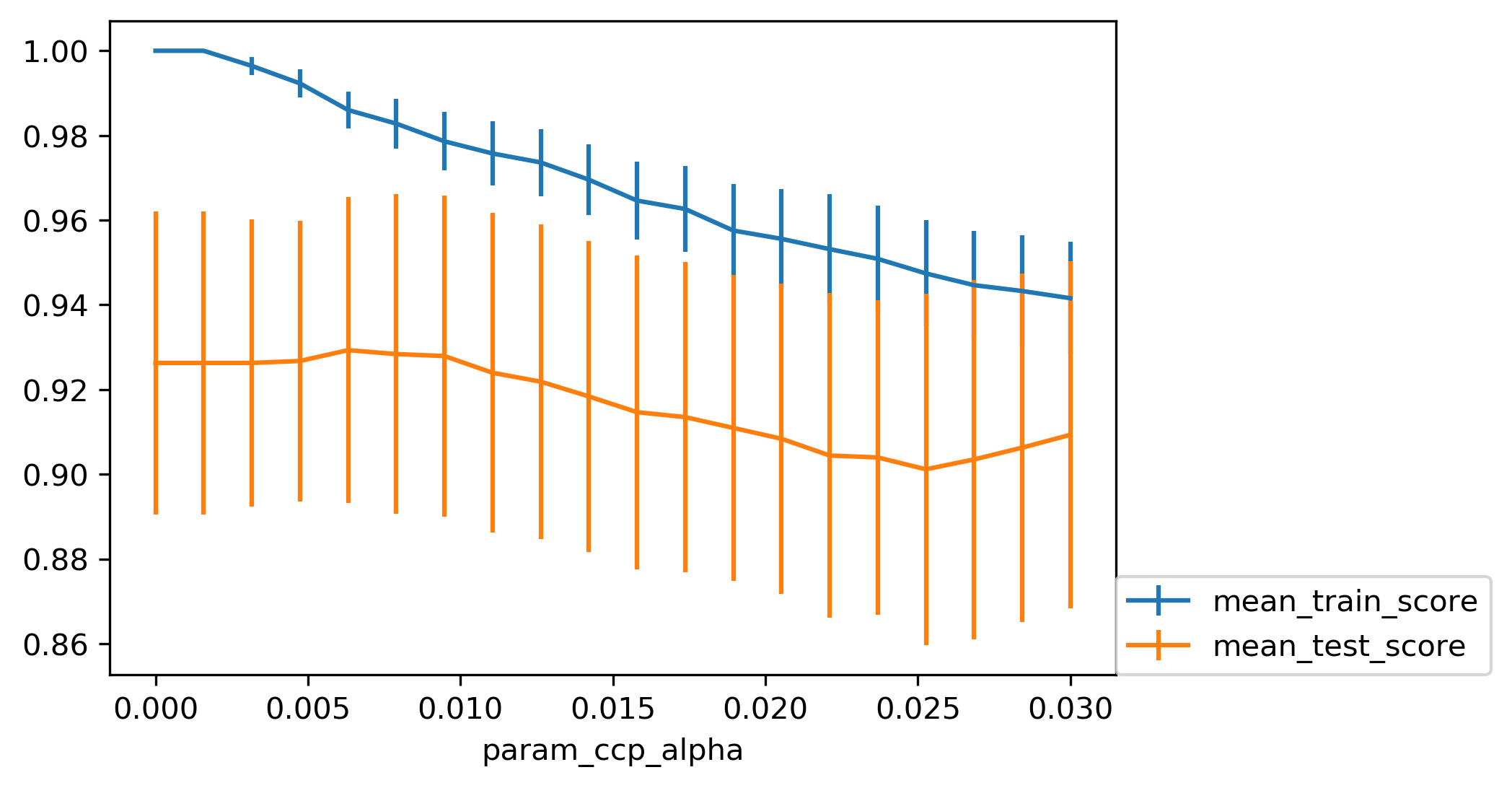
Cost Complexity Pruning
- Objective: \(R_\alpha(T) = R(T) + \alpha |T|\)
- \(R(T)\) = total leaf impurity; \(|T|\) = number of leaves; tune \(\alpha\)
Efficient Pruning Path
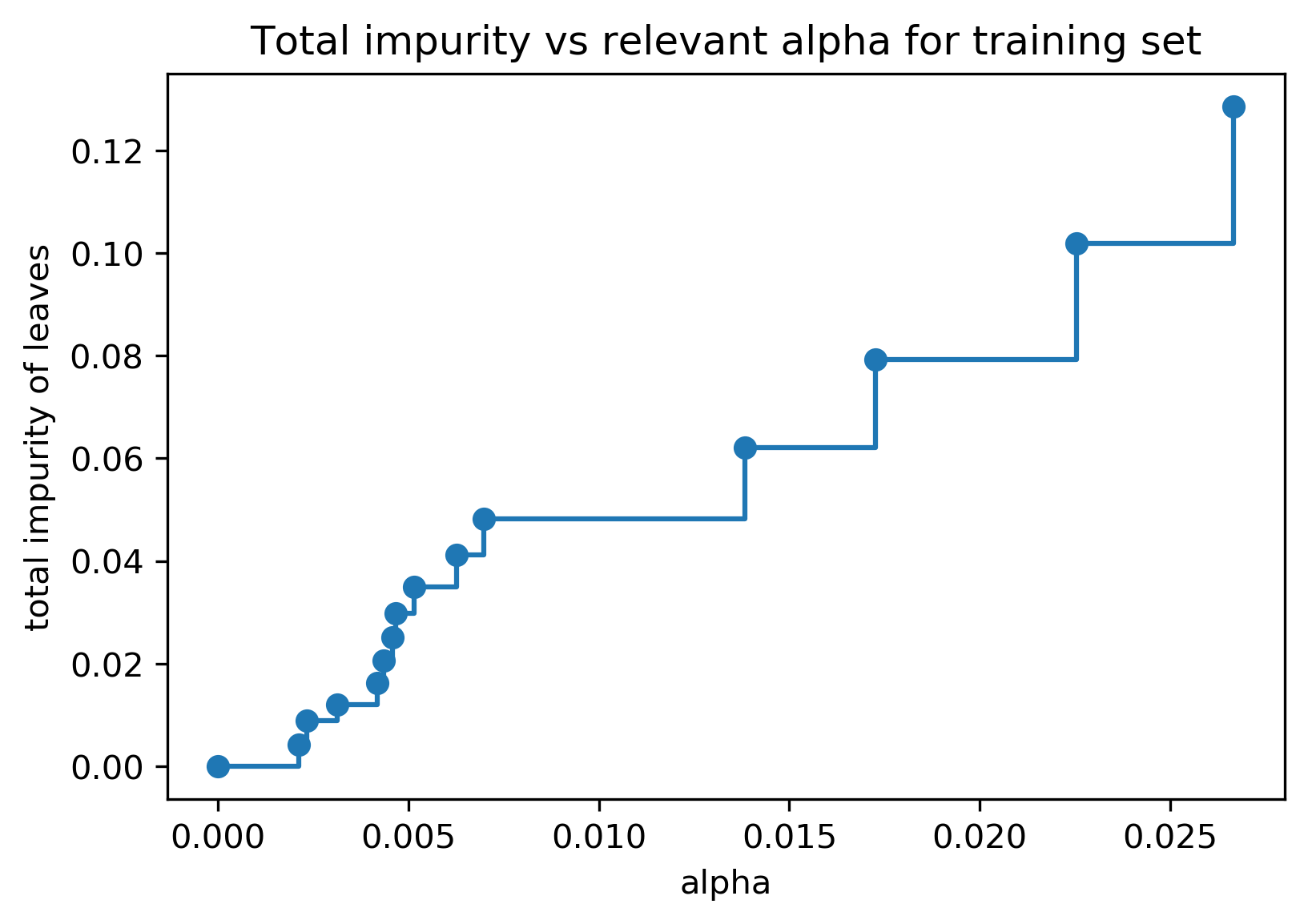
Post- vs Pre-Pruning
- Cost-complexity pruning result
- max_leaf_nodes search result

Feature Importance
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
iris.data, iris.target, stratify=iris.target, random_state=0)
tree = DecisionTreeClassifier(max_leaf_nodes=6).fit(X_train, y_train)
tree.feature_importances_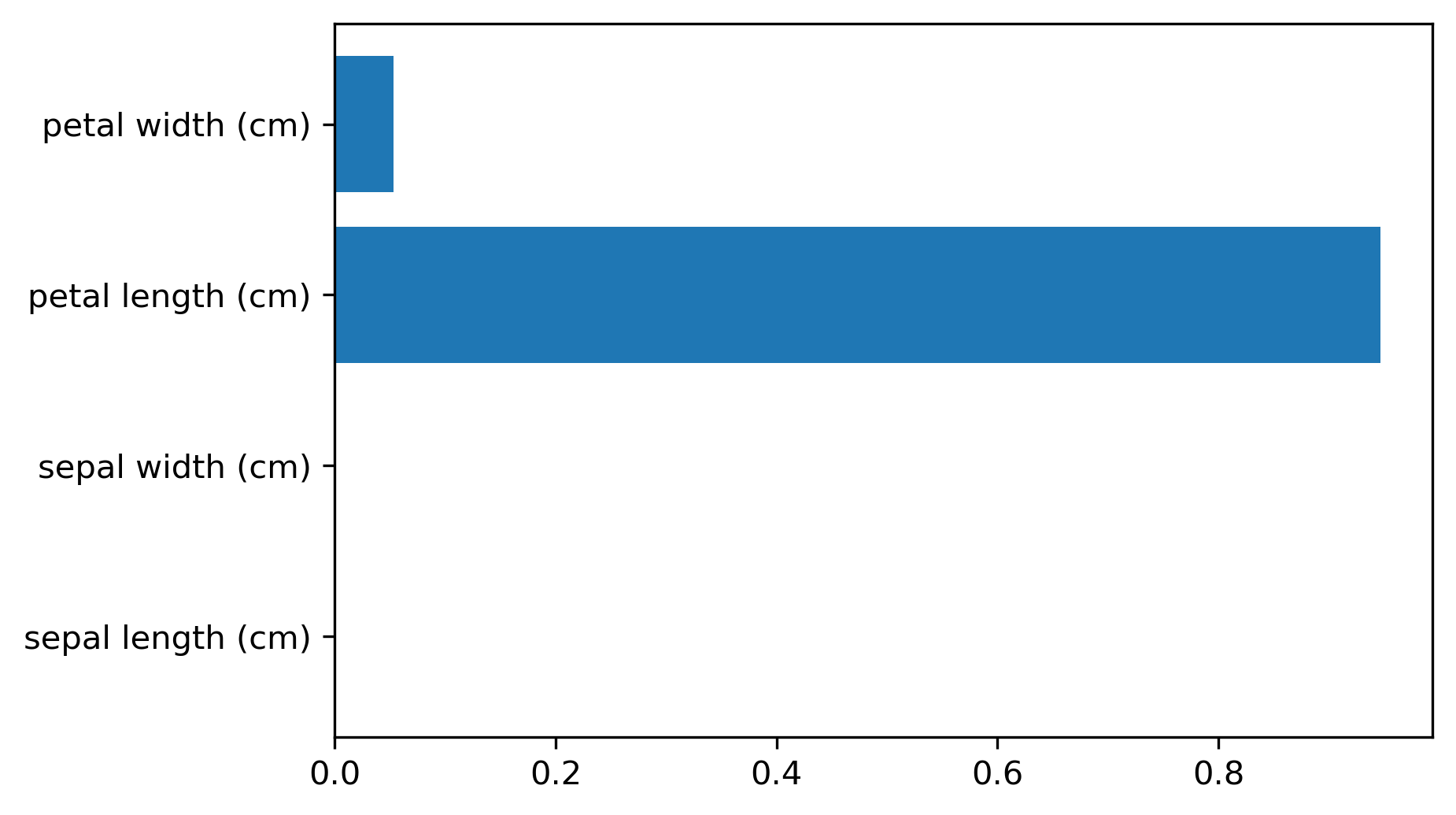
- Sum of impurity decreases per feature; magnitude only (no sign)
- Unstable with correlated features or different splits
Categorical Data
- Trees can split categories into subsets; many possible splits
- Exact search costly; efficient for binary classification + Gini
- In sklearn, one-hot encoding is needed today as sklearn can’t handle categorical features natively
Predicting Probabilities
- Leaf probability = class fraction in leaf
- e.g. for three-class leaf with samples of (AA BBB CCCCC): \(P(A)=0.2\), \(P(B)=0.3\), \(P(C)=0.5\)
- Deep, unpruned trees give overconfident (100%) probabilities
- Pruning helps but calibration may still be poor
Limitations of Trees
- Instability & overfitting: small data changes can alter splits.
- (Emsembles help)
Instability
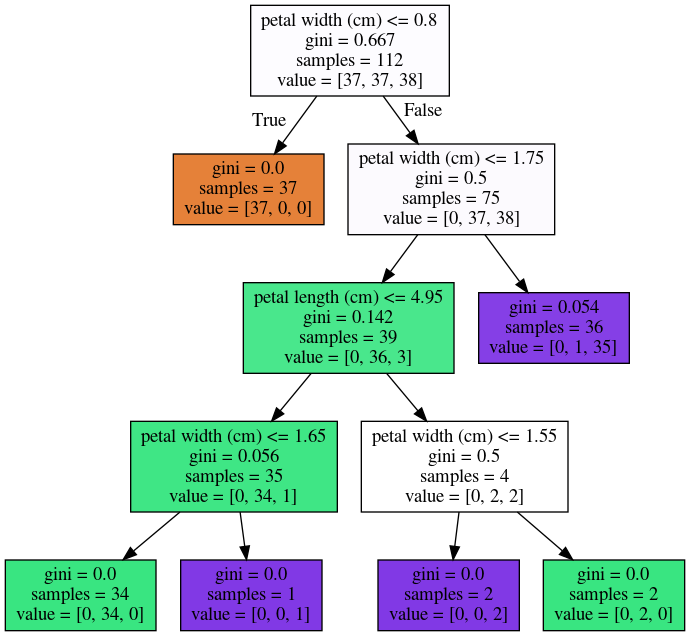
- Small data changes can alter splits
Emsemble Methods
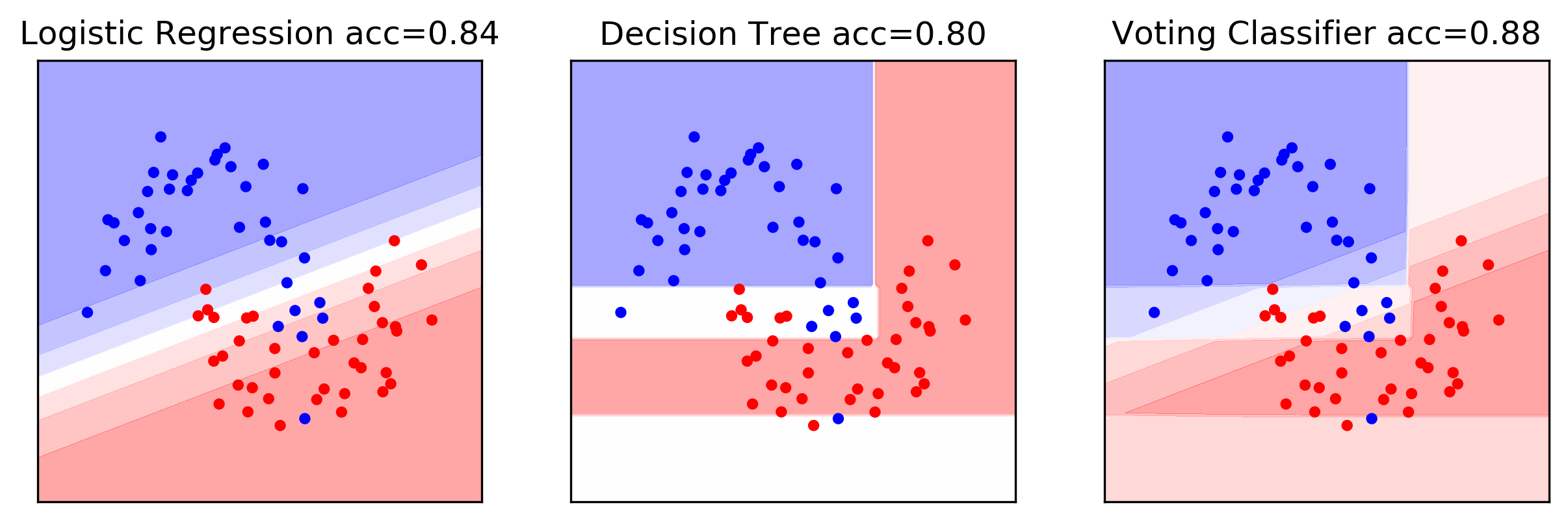
Poor man’s ensemble
- Combine multiple models to reduce variance / improve accuracy
- Train several models with different seeds; average predictions
- Owen Zhang (long time Kaggle 1st): build XGBoosting models with different random seeds.
- Works across model families (e.g., tree + linear + RF + NN)
- Key to success:
diversity among models - sklearn:
VotingClassifiersoft: average the probabilities of all models and take the class with largest average probabilityhard: let each model make a prediction and take majority vote
VotingClassifier Example
Tree ensemble: two types
- Bagging (Bootstrap Aggregation):
random forests - Boosting: gradient boosting machines (GBM),
XGBoost, LightGBM, CatBoost, etc.
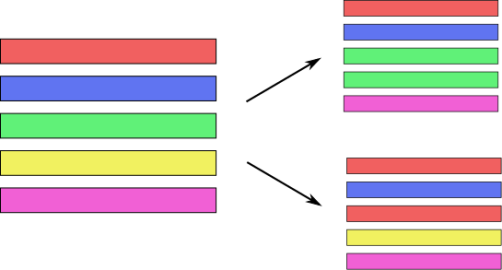
Bagging (Bootstrap Aggregation)
- Sample with replacement (same size as dataset)
- Train a model on each bootstrap sample
- Average predictions to cut variance
Bias and Variance
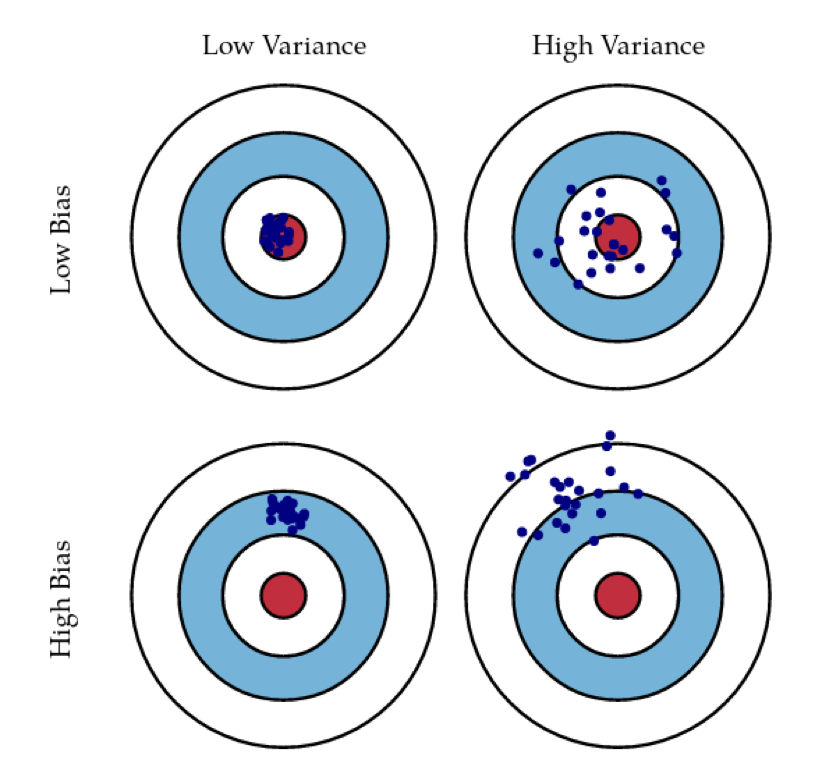
- Aim for low bias + low variance
- Averaging high-variance models can lower variance
Ensemble: Bias vs Variance
- Generalisation improves with strong base learners and low correlation
- Diversifying models (or data/features) helps more than sheer count
Random Forests
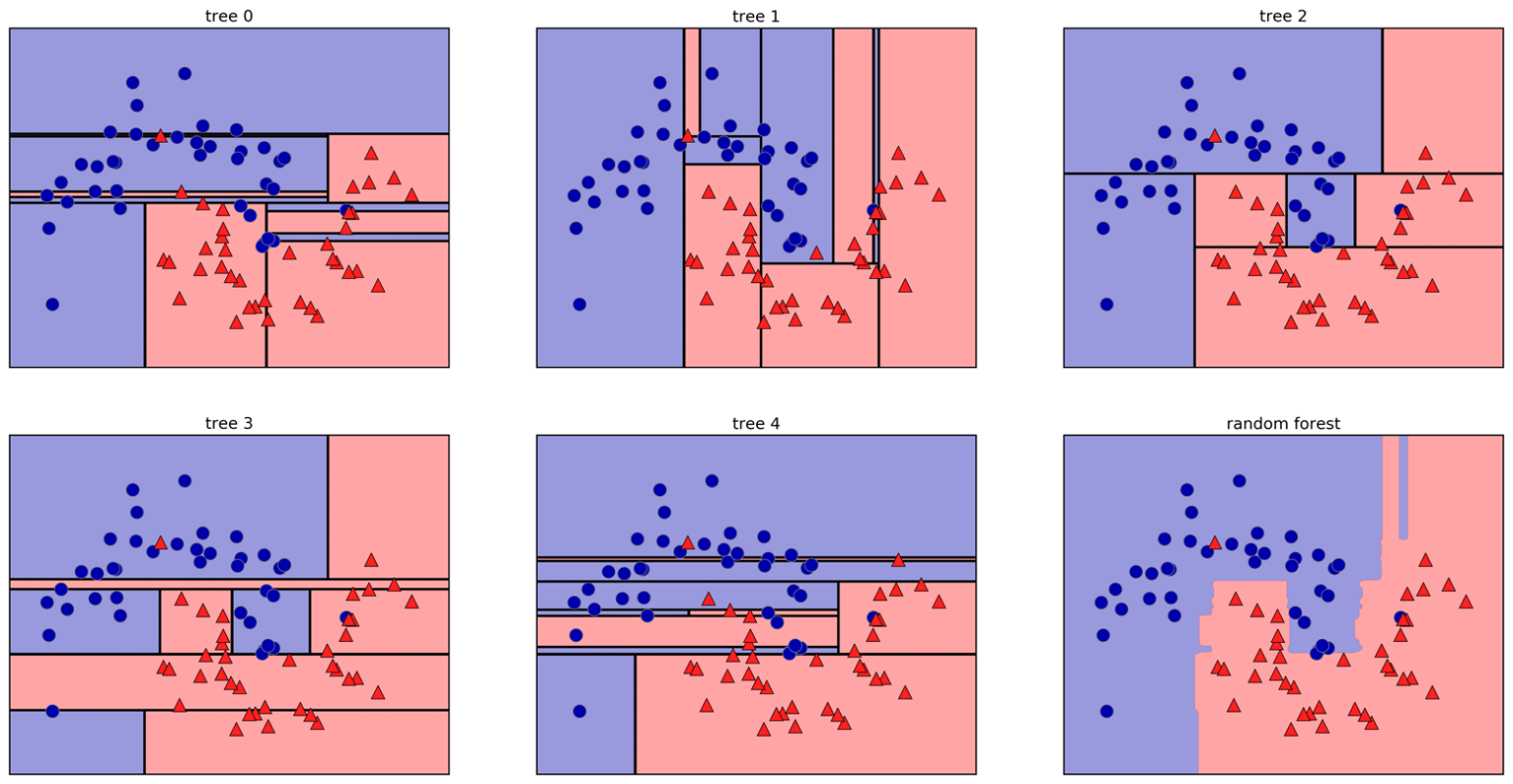
- Bagging + feature subsampling at each split
Randomise in Two Ways
- For each tree: bootstrap sample of rows
- For each split: sample features without replacement
- More trees → lower variance (diminishing returns)

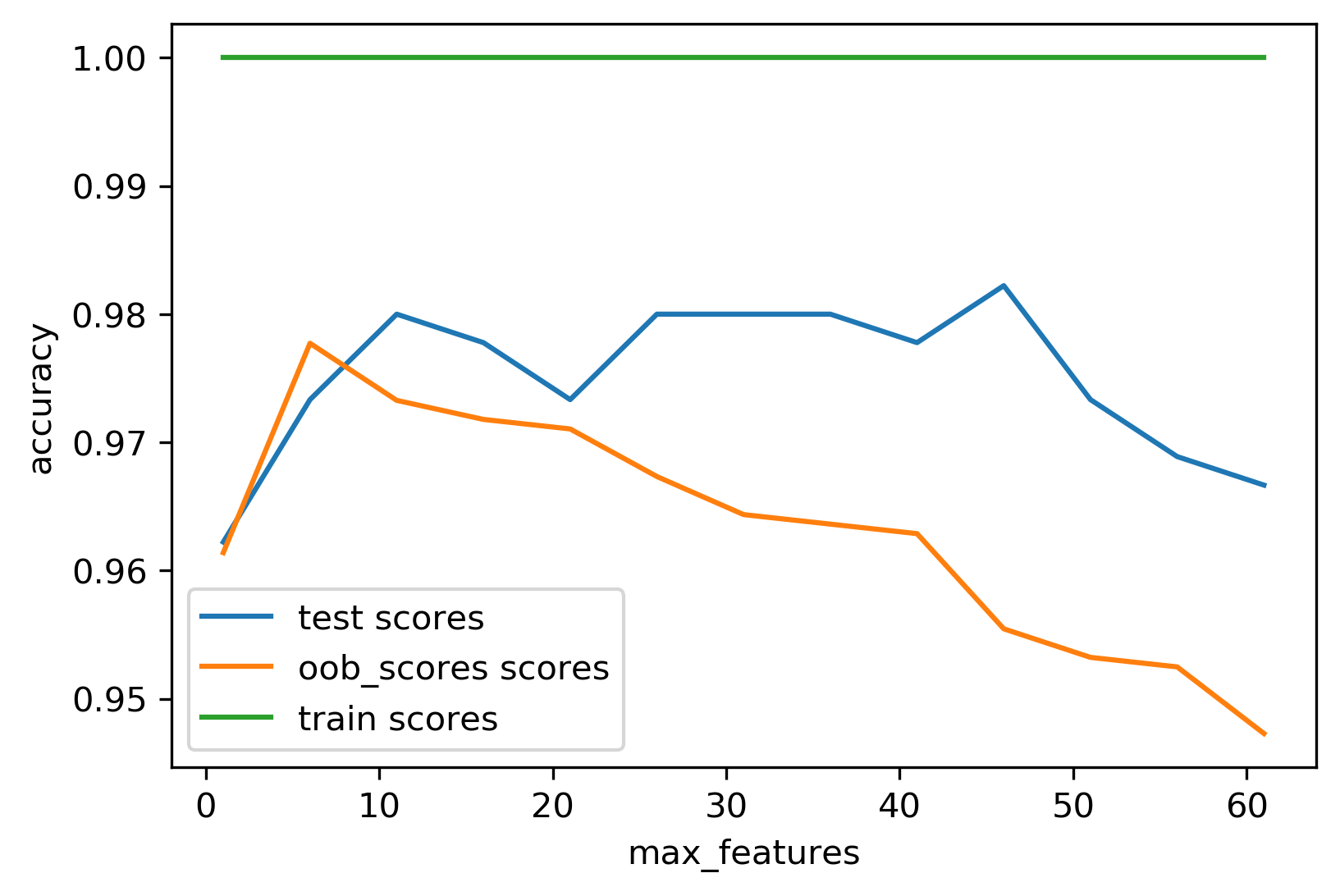
Tuning Random Forests
max_features: ~\(\sqrt{p}\) for classification, ~\(p\) for regressionn_estimators: use 100+; more reduces variance- Pre-pruning (
max_depth,max_leaf_nodes,min_samples_split) can cut size/time
Extremely Randomized Trees
- Add randomness: draw split thresholds at random per feature
- Often no bootstrap; faster (no threshold search)
- Can yield smoother boundaries; less common than standard RF
Warm-Starts
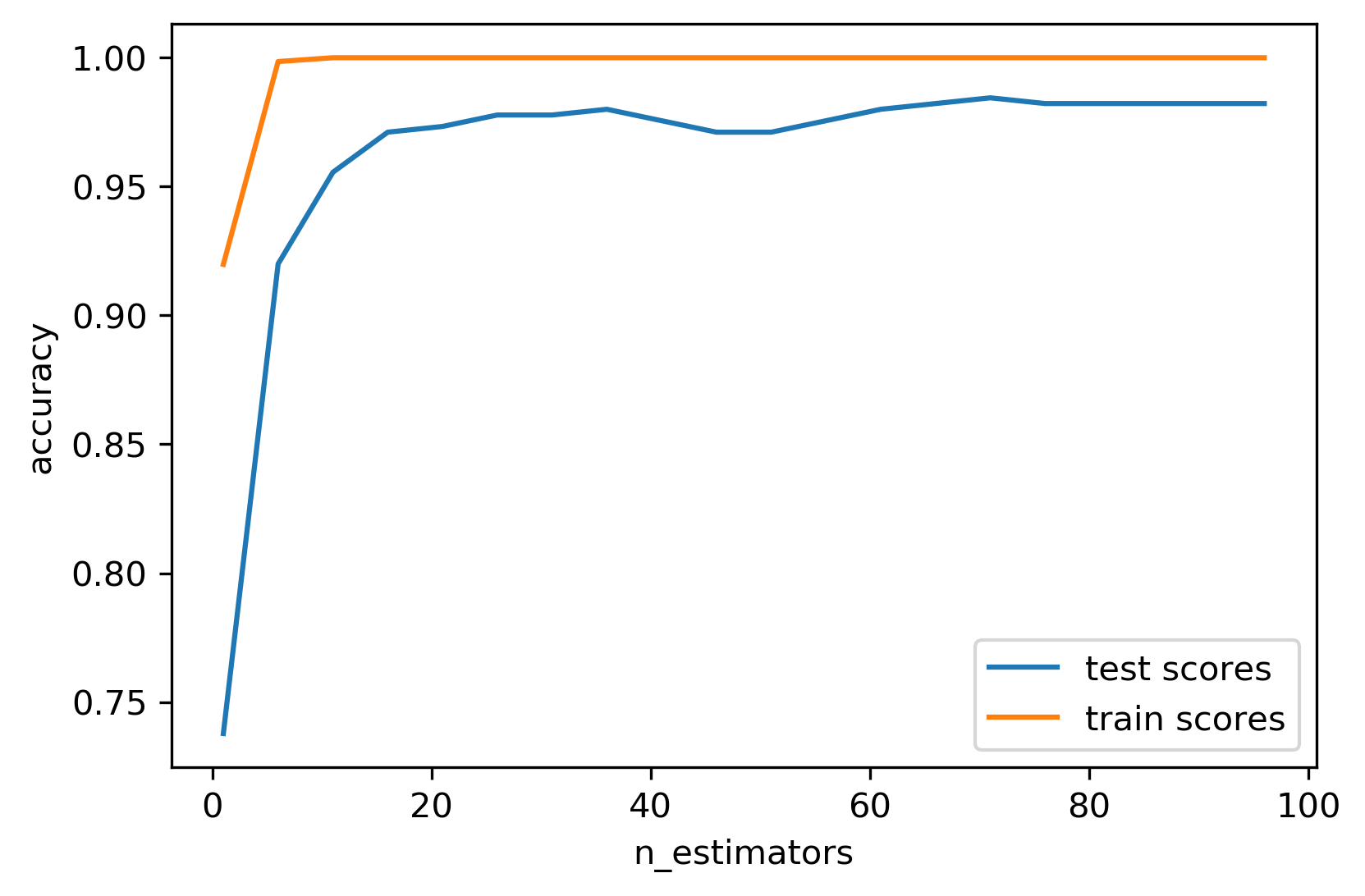
- Increase trees incrementally; stop when scores stabilise
Out-of-Bag Estimates (optional)
- Each tree trains on ~66% of data; predict remaining ~34%
- Average OOB predictions as a free validation score
Variable Importance (RF)
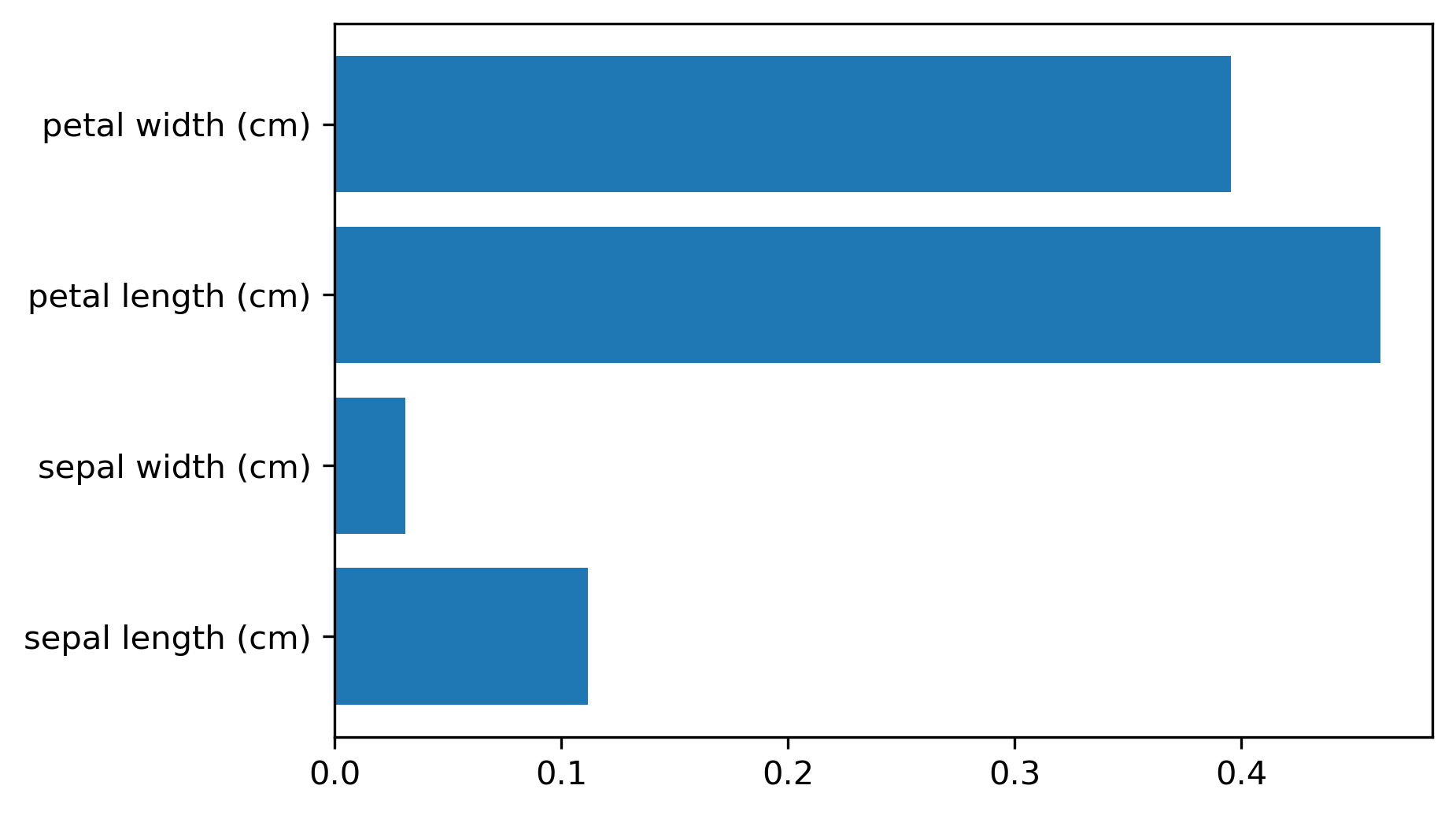
- More stable than single-tree importances; still magnitude-only
Trees: Takeaways
- Non-linear without heavy preprocessing
- Single small trees interpretable; forests robust baselines
- Beware extrapolation limits and instability
Gradient Boosting
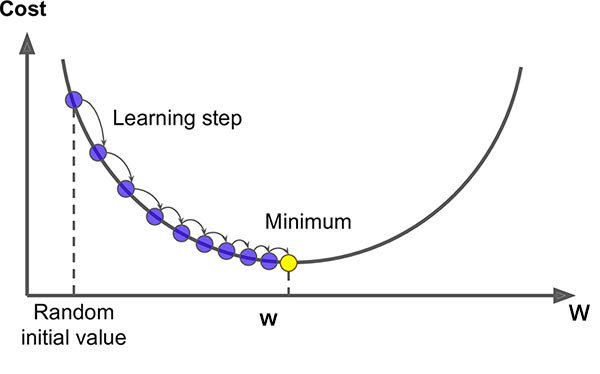
Gradient Descent
- Optimise \(\arg\min_w F(w)\) by stepping along \(-\nabla F(w)\)
- Update: \(w_{i+1} = w_i - \eta_i \nabla F(w_i)\)
- Converges to a local minimum (global for convex losses)
Stochastic Gradient Descent
- Logistic regression objective: log-loss + regularizer
- SGD uses one (or a mini-batch of) example(s) per step to approximate the gradient
- Faster on large data; noisier updates
Gradient Boosting Idea
- Train a small tree to predict \(y\)
- Train next tree on residuals of previous model (or on points poorly predicted)
- Repeat; sum scaled predictions: \(f(x)=\sum \gamma g_k(x)\) with learning rate \(\gamma\)
Gradient Boosting Algorithm
\[ \begin{aligned} f_{1}(x) &\approx y \\ f_{2}(x) &\approx y - \gamma f_{1}(x) \\ f_{3}(x) &\approx y - \gamma f_{1}(x) - \gamma f_{2}(x) \end{aligned} \]
- Each new tree fixes remaining error
- Small \(\gamma\) (e.g., 0.1) for smoother learning
Gradient Boosting as Gradient Descent
- Linear regression minimizes \(\sum (y - w^T x - b)^2\)
- Gradient descent updates weights
- Gradient boosting minimises same loss over predictions \(\hat{y}\)
- Update \(\hat{y}\) by adding trees along negative gradient
Regression Example
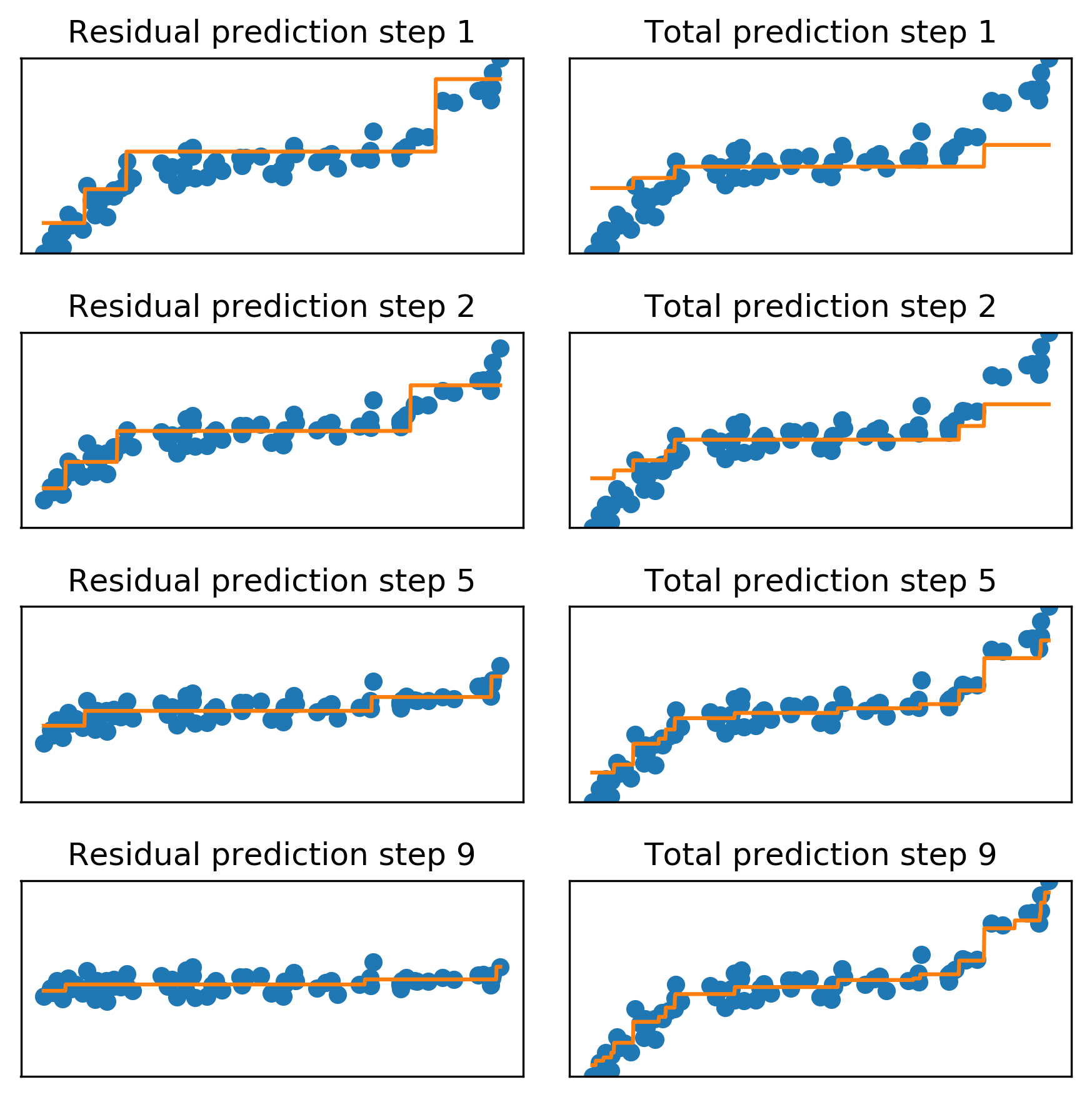
- Shallow trees fit residuals sequentially until residuals shrink
Classification Example
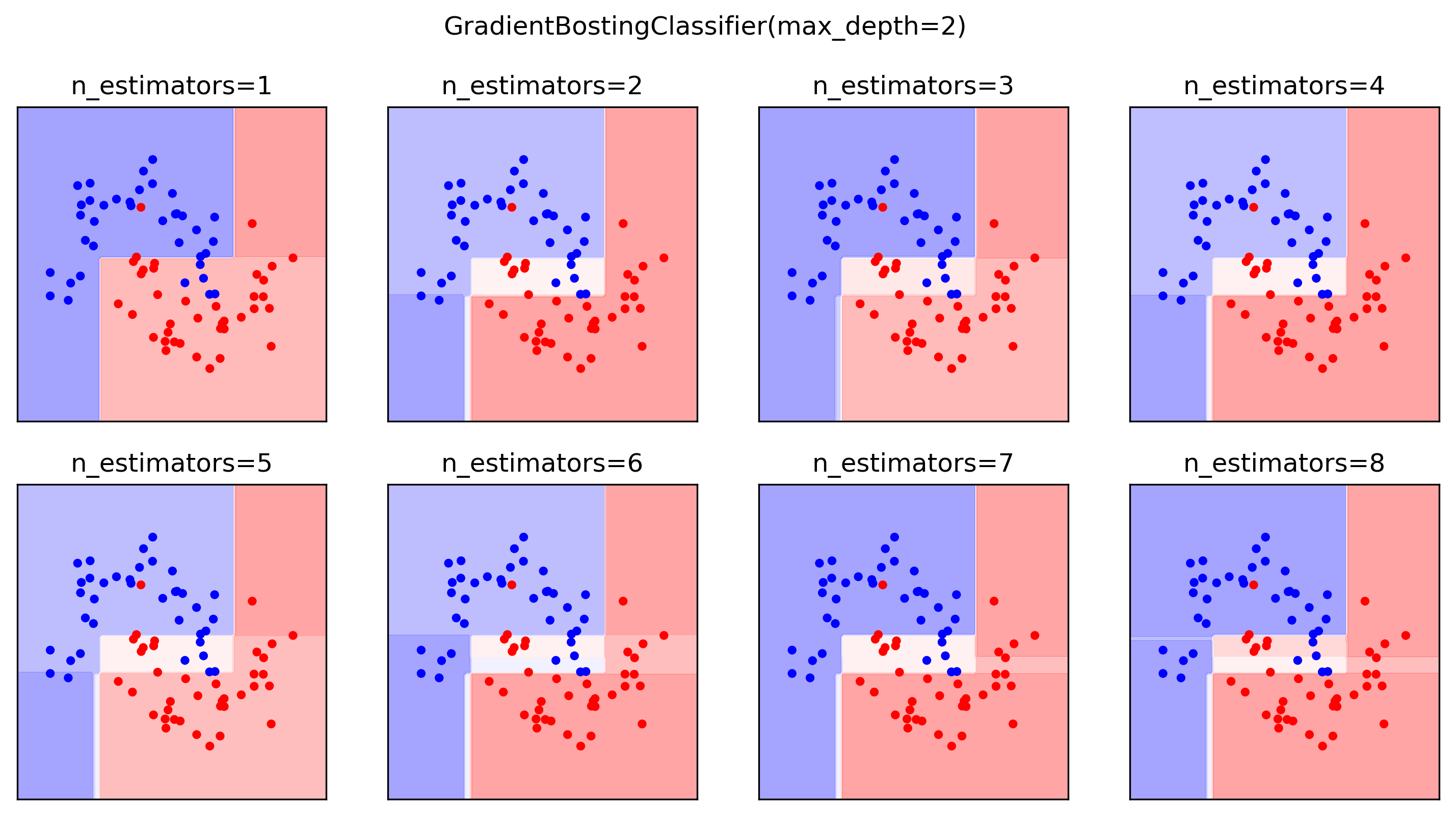
- Probability surfaces become sharper as trees accumulate
- Multiclass: one regression tree per class per step
Early Stopping
- More trees can overfit
- Stop when validation metric stops improving
- Either fix \(n_{\text{estimators}}\) and tune \(\gamma\), or fix \(\gamma\) and stop early
Tuning Gradient Boosting
- Use shallow trees (strong pruning via
max_depth) - Tune learning rate,
n_estimators, subsampling of rows/columns - Optional regularisation (
min_samples_split,min_impurity_decrease)
Ensuring uncorrelated trees: Aggressive Subsampling
- Row and column subsampling reduce correlation and overfitting
- Common in XGBoost/LightGBM
XGBoost
- Fast, supports missing values, GPU/cluster training
- L1/L2 on leaves; fast approximate splits; sparse data friendly
LightGBM
- Native categorical handling; missing values; GPU/cluster support
CatBoost
- Strong on categorical data; symmetric trees; target-encoding tricks; GPU
Advantages of Gradient Boosting
- Often more accurate than random forests with tuning
- Small models; fast prediction
- Hist/XGB/LightGBM implementations are very fast
Summary: when to use a tree or tree ensemble
- For tabular data
- Can model non-linear relationships and require minimal preprocessing
- Single small tree for interpretability, but a single tree often underfits and doesn’t generalise well
- Random forests are very robust and good benchmark
- Gradient boosting generate best accuracy when tuned
Questions
© CASA | ucl.ac.uk/bartlett/casa
